OrthoInspector
What is OrthoInspector ?
OrthoInspector is a package dedicated to orthology inference and analysis. The package can be installed for local use on a desktop computer or configured for a server implementation. Its purpose is to facilitate the installation of a database of newly inferred orthology relations and facilitate the exploitation of the resulting orthology database by both expert and non-expert users.
What are the requirements ?
OrthoInspector is compatible with all systems supporting Java 1.6.x JVM and superior. Its components are based on a SQL database / Java client architecture. Consequently, you need to setup an access to an SQL engine (locally or remotely). Any database engine can be used as soon as a Java JDBC driver is available for it (see complete Requirements for details). Exploiting the inferred orthologs will also require a local installation of the latest NCBI blast+ package.
Why do I need a SQL database ?
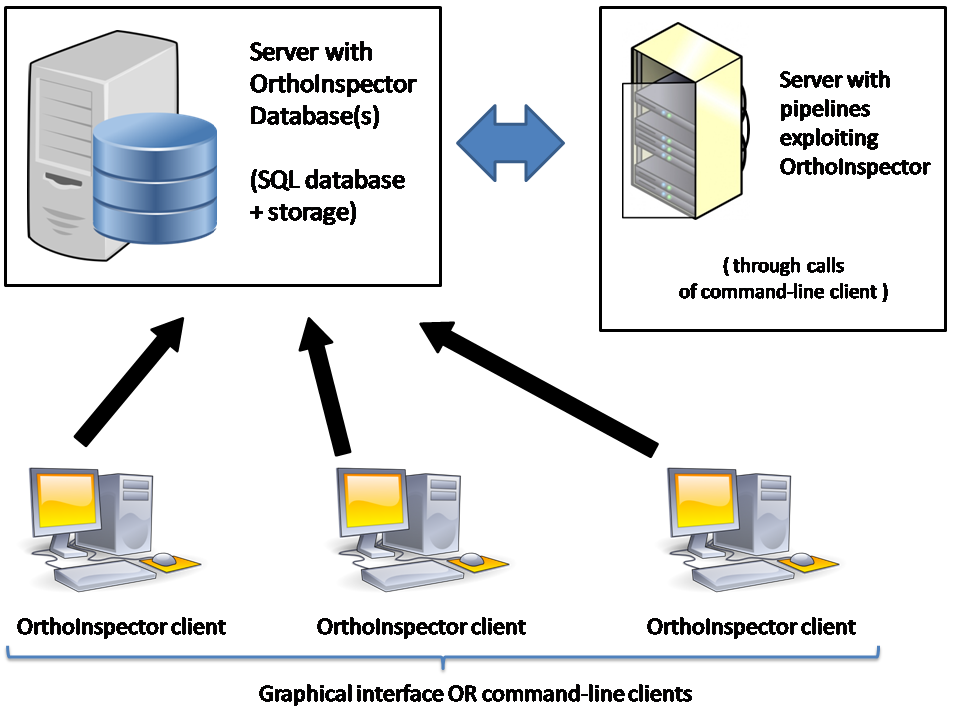
While the package can be installed on a single machine, we choose to code OrthoInspector in Java and through a SQL database / Java client architecture to facilitate its exploitation in a network environment. Orthology relations are first predicted from a set of proteomes and a blast all-against-all. They are transferred to an SQL database and can be exploited by several clients, allowing several users to query the database via a graphical interface or allowing several pipelines to exploit these data simultaneously.
Be aware that OrthoInspector can be installed either on a network or on a single desktop computer ! You just need to setup an accessible SQL engine, locally (desktop computer) or remotely (office network). The network configuration described below is a recommendation but can be adapted to your needs.
How is organized the package and what can I do with it ?
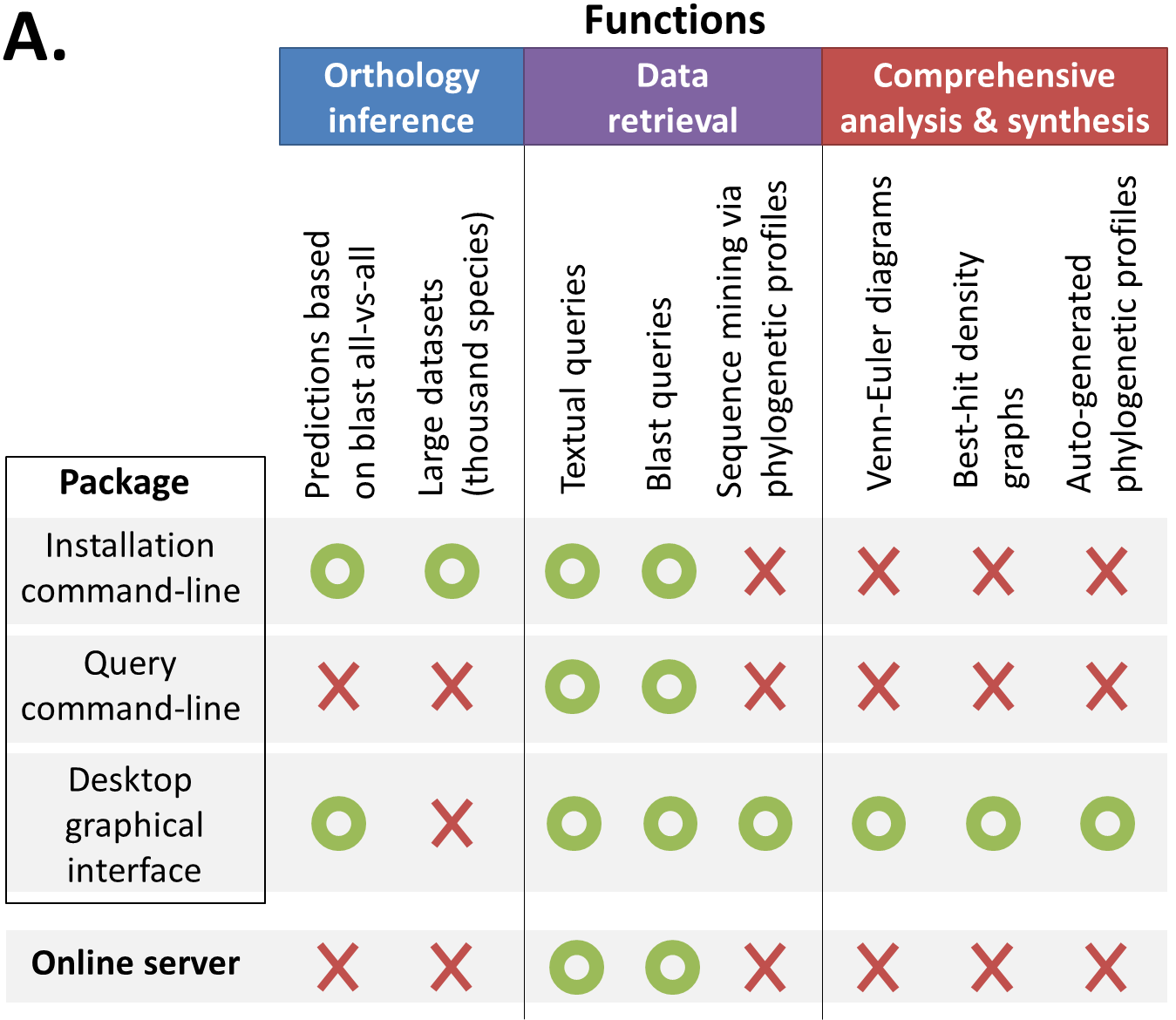
The package is separated in three Java executables (and a few configuration files) :
Installation command-line: A command-line to predict orthology relations and transfer them in a SQL database.
Query command-line: A command-line to query these predictions or export them from the database.
Graphical interface: a desktop interface which regroup most high-level functionnalities (complex queries and visualisation tools) but can also be used directly to infer small orthology datasets (<100 species).
Both the Installation command-line and the Graphical interface can be used to infer orthology relations. The query command-line and most other functions from the graphical interface are designed to exploit these predictions.
Installation command-line:
This component is recommended to infer large orthology databases from large datasets (> 100 species). A similar procedure is available in the graphical interface but the Installation command line provides several useful options to handle larger datasets. For instance, it can read compressed blast inputs, create compressed outputs, share predictions to several CPUs and facilitate crash recovery (useful when your calculations require a weeks of CPU...).
Query command-line:
This component is designed for basic data retrieval from the orthology database. You can query the databases with textual searches, fasta queries or export the entire set of orthologs related to a particular organism. It can easily be integrated into other bioinformatic pipelines.
Graphical interface:
Contains most functionalities and is a good option for non-specialists which want to avoid command-line manipulations (but you still need to install and configure the SQL database). You can install a small database (<100 organisms) with an automated procedure. This procedure is fully automated if you use MySQL or Postgresql engines. The graphical interface contains advanced querying tools, such as sequence extractions based on phylogenetic profiles. All the graphical tools dedicated to orthology analysis and visualization (Euler diagrams, automated phylogenetic profile construction...) are also in the Graphical interface.
To know more about these tools, consult the dedicated tutorials which are available on this website.
What about the online databases ?
Please check out the page describing our available databases.
Licensing:
OrthoInspector is licensed under the GNU GPL version 3. You can download it, modify it and share it at your convenience but don't forget to cite the authors ! ;-)